suppressPackageStartupMessages(library(cmdstanr))
suppressPackageStartupMessages(library(magrittr))
suppressPackageStartupMessages(library(tidybayes))
suppressPackageStartupMessages(library(ggplot2))Grouped data
Outline
Topics
- Using the package
tidybayesto feed complex data into Stan.
Rationale
Data organized in various groups is a frequent feature of Bayesian models, in particular in hierarchical models.
Getting grouped data into stan is tedious and error-prone. The package tidybayes automates much of this.
Pre-reading
If you have never heard about “tidy data”, while it is not strictly essential for this course, it is a good investment to skim this tutorial on tidy data.
Example
First, install the packages magrittr and tidybayes (and ggplot2 if you have not done so already), then import them:
Data prep
We load data modelling covid vaccines efficacy:
data = read.csv(url("https://raw.githubusercontent.com/UBC-Stat-ML/web447/main/exercises/ex06_assets/vaccines_full.csv"))
data$is_vaccinated = ifelse(data$arms == "vaccinated", 1, 0)
rmarkdown::paged_table(data)The magic conversion of “tidy data” (data in a format like the above) into a format that can be consumed by Stan is done using compose_data:
stan_converted = compose_data(data)
stan_converted$trials
[1] 1 1 3 3 2 2
$n_trials
[1] 3
$arms
[1] 2 1 2 1 2 1
$n_arms
[1] 2
$groupSizes
[1] 5807 5829 18198 18325 14134 14073
$numbersOfCases
[1] 30 101 8 162 11 185
$is_vaccinated
[1] 1 0 1 0 1 0
$n
[1] 6Stan model
Here we consider a simple, non-hierarchical model but fitting all the data at once. As an exercise, think about how you can modify this model to be hierarchical.
vaccines.stan
data {
int n;
int n_trials;
array[n] int<lower=1,upper=n_trials> trials;
array[n] int arms;
int n_arms;
array[n] int groupSizes;
array[n] int numbersOfCases;
array[n] int is_vaccinated;
}
parameters {
vector<lower=0,upper=1>[n_trials] efficiencies;
vector<lower=0,upper=1>[n_trials] prevalences;
}
model {
for (trial in 1:n_trials) {
efficiencies[trial] ~ beta(1, 1);
prevalences[trial] ~ beta(1, 1);
}
for (i in 1:n) {
numbersOfCases[i] ~ binomial(groupSizes[i], prevalences[trials[i]] * (is_vaccinated[i] == 1 ? 1.0 - efficiencies[trials[i]] : 1.0));
}
}Fitting
The fit object returned by sample() does not know about the string labels attached to each trial integer index. We use fit %<>% recover_types(data) to add back that information:
mod = cmdstan_model("vaccines.stan")
fit = mod$sample(
seed = 1,
data = stan_converted,
output_dir = "stan_out",
refresh = 0,
iter_sampling = 10000
)
fit %<>% recover_types(data)Getting draws
The output of Stan is not tidy either.
fit variable mean median sd mad q5 q95 rhat ess_bulk
lp__ -2793.20 -2792.87 1.74 1.58 -2796.52 -2790.99 1.00 18550
efficiencies[1] 0.69 0.69 0.06 0.06 0.57 0.78 1.00 47395
efficiencies[2] 0.93 0.94 0.02 0.02 0.90 0.96 1.00 52259
efficiencies[3] 0.94 0.95 0.02 0.02 0.91 0.97 1.00 48810
prevalences[1] 0.02 0.02 0.00 0.00 0.01 0.02 1.00 44590
prevalences[2] 0.01 0.01 0.00 0.00 0.01 0.01 1.00 51935
prevalences[3] 0.01 0.01 0.00 0.00 0.01 0.01 1.00 51276
ess_tail
26750
29969
30050
28217
30768
29448
30109Use spread_draws to put the draws into a tidy format:
fit %>% spread_draws(efficiencies[trials], prevalences[trials]) %>% head(10)# A tibble: 10 × 6
# Groups: trials [1]
trials efficiencies .chain .iteration .draw prevalences
<chr> <dbl> <int> <int> <int> <dbl>
1 AZ-Oxford (combined) 0.656 1 1 1 0.0153
2 AZ-Oxford (combined) 0.626 1 2 2 0.0169
3 AZ-Oxford (combined) 0.739 1 3 3 0.0168
4 AZ-Oxford (combined) 0.731 1 4 4 0.0160
5 AZ-Oxford (combined) 0.768 1 5 5 0.0189
6 AZ-Oxford (combined) 0.552 1 6 6 0.0158
7 AZ-Oxford (combined) 0.608 1 7 7 0.0164
8 AZ-Oxford (combined) 0.695 1 8 8 0.0188
9 AZ-Oxford (combined) 0.809 1 9 9 0.0195
10 AZ-Oxford (combined) 0.755 1 10 10 0.0184Plotting
Now that we have draws in tidy format, instead of using specialized MCMC plotting libraries we can just use ggplot:
fit %>%
spread_draws(efficiencies[trials]) %>%
ggplot(aes(x = efficiencies, y = trials)) +
stat_halfeye() +
theme_minimal()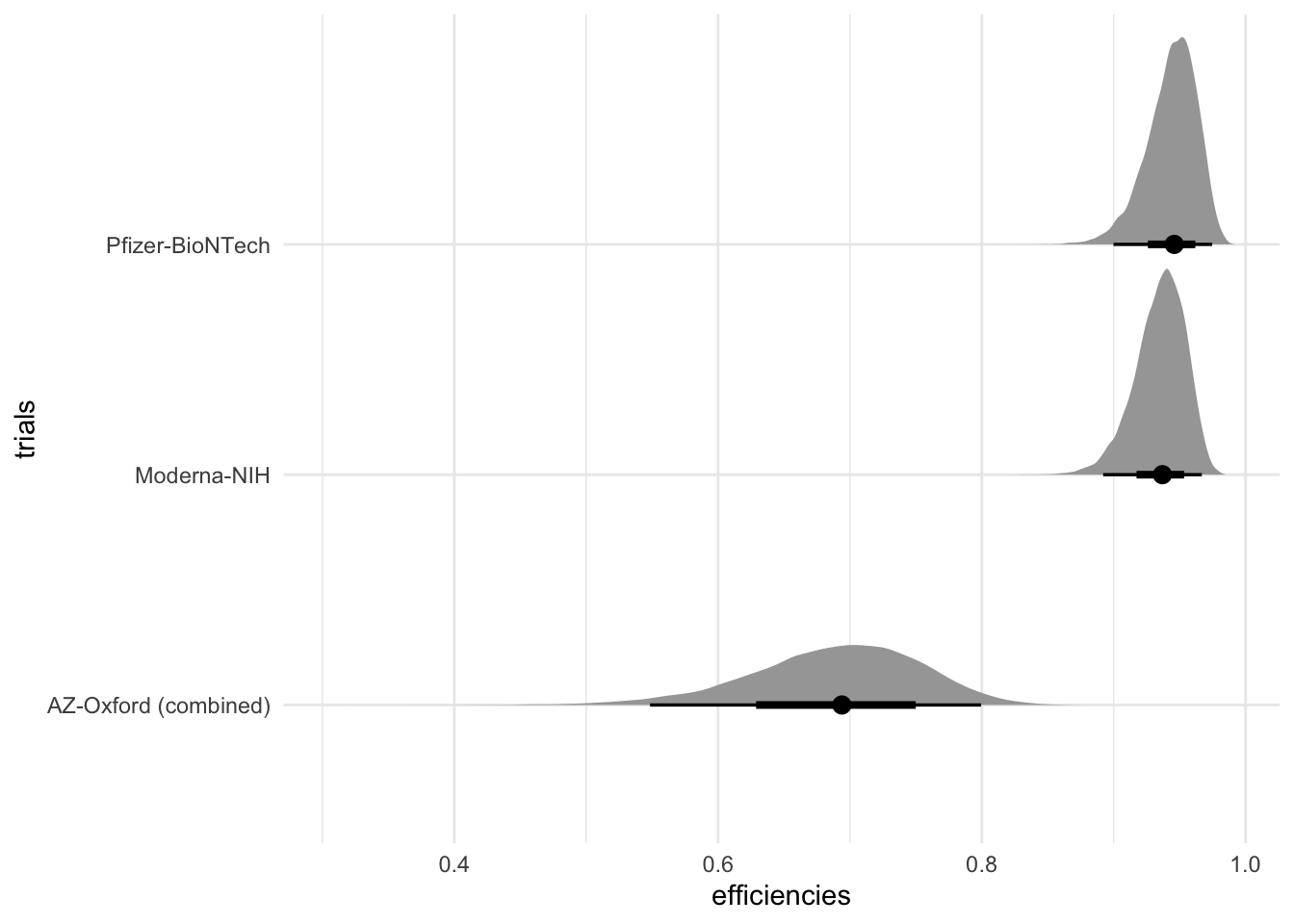
This makes it easier to customize plots.
Summaries
tidybayes also offers convenient ways to compute summaries such as High Density Intervals (HDI)
fit %>%
spread_draws(efficiencies[trials]) %>%
median_hdi(efficiencies)# A tibble: 3 × 7
trials efficiencies .lower .upper .width .point .interval
<chr> <dbl> <dbl> <dbl> <dbl> <chr> <chr>
1 AZ-Oxford (combined) 0.694 0.558 0.806 0.95 median hdi
2 Moderna-NIH 0.937 0.897 0.970 0.95 median hdi
3 Pfizer-BioNTech 0.946 0.905 0.978 0.95 median hdi More information
See the tidybayes documentation.